The intricate quest to discover where tetracyclines go in human cells
By Caitlin McDermott-Murphy
We know that antibiotics treat bacterial infections. We also know why they work. Tetracycline antibiotics, for example, stop bacteria from making protein. Like a boot on a wheel, the drugs bind to the bacterial cell’s ribosome—where protein is made—and prevent it from working. Without protein, the bacteria weaken and die.
Recently, researchers discovered a possible new job for these tetracycline antibiotics: treatment for pathological inflammation and cancer in humans.
This treatment has proven effective in clinical trials. Two patients, a young man and an elderly woman[1], who suffered from rare, stubborn diseases experienced complete remission after treatment with doxycycline, a type of tetracycline. Other tetracycline varieties have shown potential in early clinical trials with patients who have a number of diseases, including refractory metastatic cancers, rheumatoid arthritis and osteoarthritis, fragile X syndrome, AIDS-related Kaposi’s sarcoma, and abdominal aortic aneurysm.
And yet, despite these promising results, exactly how the treatment works remained elusive.
|
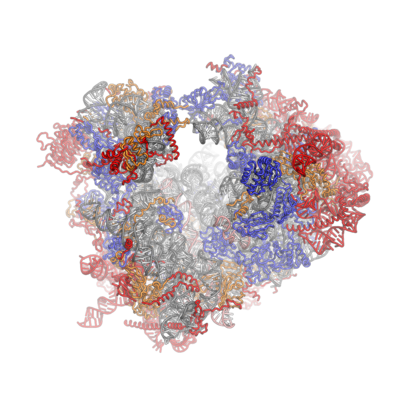
The structure of a eukaryotic (human) ribosome. Ribosomes are responsible for protein synthesis in our cells. |
Now, in a new paper published in Cell Chemical Biology, Andrew Myers and his team provide an answer. “Despite these promising clinical results and a wealth of published scientific research,” the paper states, “no relevant target(s) or mechanism(s) have been clearly identified to account for these non-canonical effects of tetracyclines.” This knowledge gap impedes further drug development and treatment.
So, Myers and his team set out to track and map exactly what happens when two tetracycline analogs, doxycycline and Col-3, enter a cell. This is far from easy. Like tracking a mouse in a blizzard, “the detective work to find targets of drug molecules is painstakingly difficult,” says Jon Mortison, the paper’s first author.
The team used a powerful technique (“affinity isolation in tandem with mass spectrometry (MS)-based quantitative SILAC—stable isotope labeling by amino acids in cell culture— proteomics”) to isolate each drug’s target. In so doing, they observed that both analogs target our cell’s ribosome, just as they do in a bacterial cell.
“To our knowledge,” the paper notes, “this represents the first direct evidence that tetracyclines have observable effects on human cytosolic ribosomes.”
To uncover this evidence, the team used a novel strategy to hunt down the tetracycline targets. They believe their method could be adopted by other researchers for similar purposes, to build a broader map for how drugs interact with our bodies. “RNA targets in drug discovery is becoming an area of increasing interest,” Mortison said. “Some large companies and biotechs are already jumping into the space of identifying RNA targets for their pipelines.”
Going forward, the team hopes to clarify whether the exact point where doxycycline and Col-3 bind to our ribosomes could boost their ability to slow the spread of inflammation and cancer. Such precise information would provide crucial insights for drug development.
Further exploration is required to compile this intel. But Myers and his team have provided a crucial foundation for these studies to proceed. Soon, tetracyclines could be a valuable treatment option for numerous diseases, including cancer. In the meantime, the team’s new technique could, like a well-designed field guide, hasten drug discovery for myriad diseases.